Getting unique values from columns: performance comparison
Ed Baker
2024-10-21
Introduction
Getting a set of unique values from a column is a common operation in
data analysis. A commonly suggested technique is to use the
sort and uniq commands in combination. This
study compares the performance of this technique with some variations on
a real-world dataset and some contrived test examples.
Method
Datasets
The dataset is two files derived from a GBIF exported with 89,732,452 rows and 15 columns (GBIF.Org User 2024). The investigation focusses on two columns from this dataset:
publisherwith 30 unique values,licensewith 175,716 unique values.
These columns were isolated into separate files for each column to ensure the test is not influenced by column separation.
In order to compare with three more theoretical benchmarks three additional files (with the same number of lines as the real world example) were created:
same: A file where every line reads “BioAcoustica: Sound Library”unique: A file with incrementing integersalternate: Alternating lines of true and false
Machine details
The tests were performed on a VMWare virtualised instance of Ubuntu 22.04.05 LTS with 12GB of RAM and 4 Intel Xeon Platinum 8380 CPUs running at 2.30GHz.
Timing
The bash time function was used to measure the time
taken to extract unique values from each column, with 10 repetitions for
each method. As the real time taken (as reported by time)
may be affected by other processes running on the system, the sum of
user and sys time were used as the
total measure of performance, although the
real time is also reported as a measure of the time taken
to run the command and assess the impact of multi-thread operations.
Tests of single core performance were not conducted as it’s 2024.
The following methods were tested:
sort -uuniq | sort -usort | uniquniq | uniq | sort -uuniq | sort | uniq
The file being tested was read by the first sort or
uniq command, and the output was piped to
/dev/null.
Notes on sort
The sort command is used to sort the input data. The
-u option is used to remove duplicate lines.
Notes on uniq
The uniq command is used to remove duplicate lines from
a sorted file. If the file is not sorted uniq will only
output the current line if it is different from the previous line.
In real-world datasets values in a column often occur in groups, so
the combinations uniq | sort | uniq and
uniq | uniq | sort -u may effectively reduce the number of
lines piped to sort.
Results
Results by file
license column
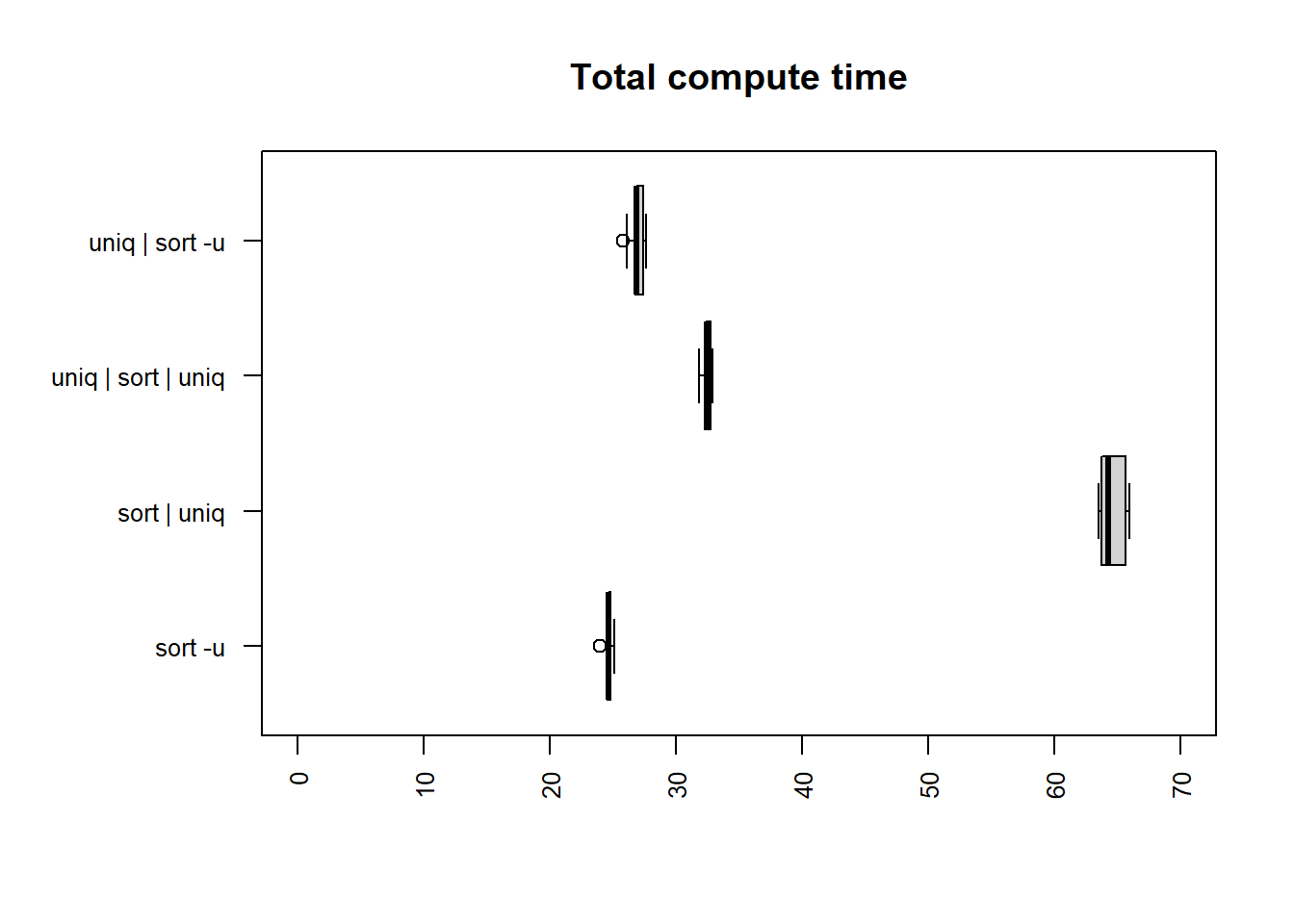
| method | mean |
|---|---|
| sort -u | 24.6963 |
| uniq | sort -u | 26.9240 |
| uniq | sort | uniq | 32.5221 |
| sort | uniq | 64.5948 |
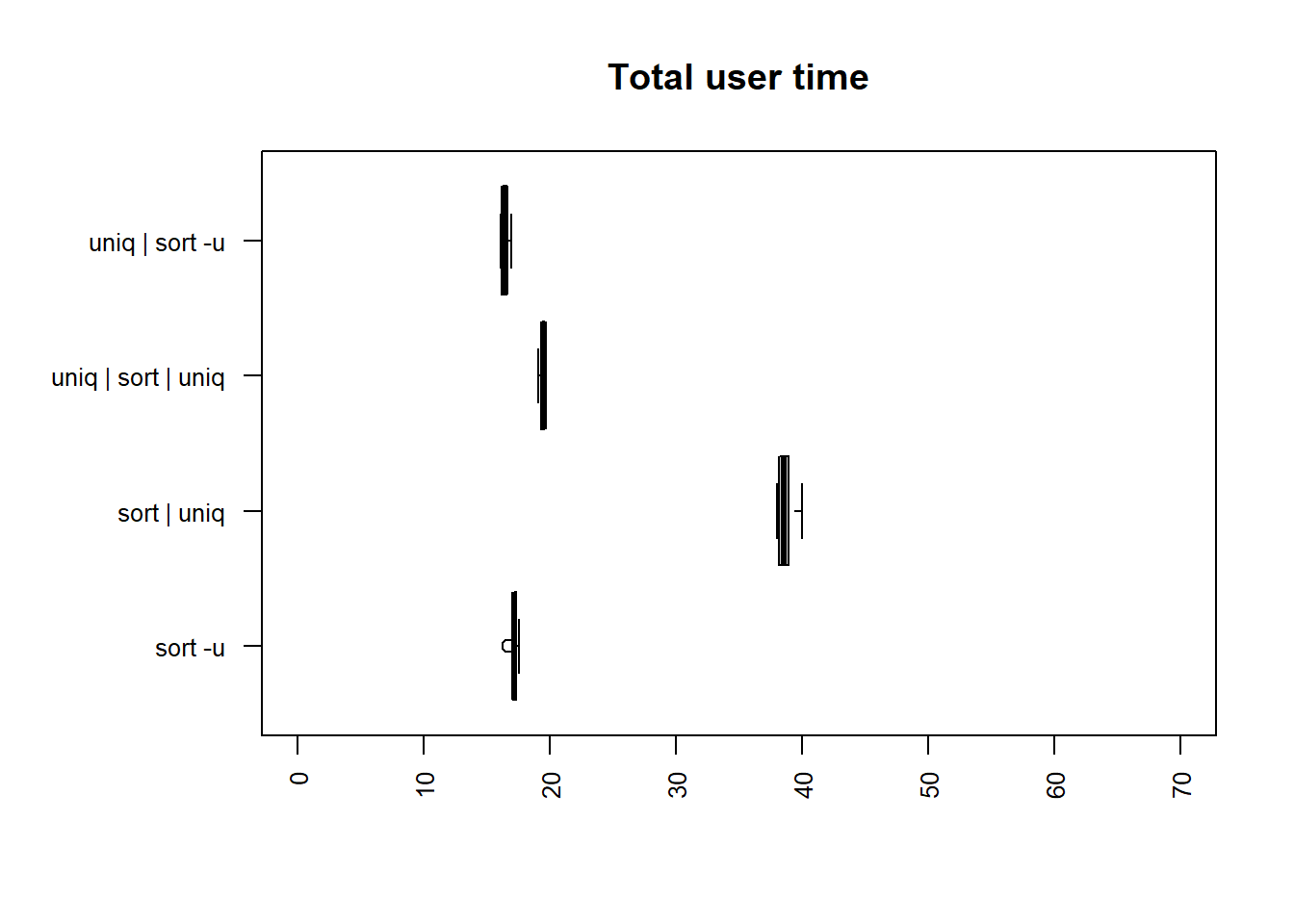
| method | mean |
|---|---|
| uniq | sort -u | 16.4382 |
| sort -u | 17.2138 |
| uniq | sort | uniq | 19.4527 |
| sort | uniq | 38.7181 |
The two fastest methods for both user and total time are
uniq | sort -u and sort -u.
sort | uniq is the slowest method, taking 2.36 times longer
than uniq | sort -u for the user, and 2.62 times longer for
the total time.
publisher column
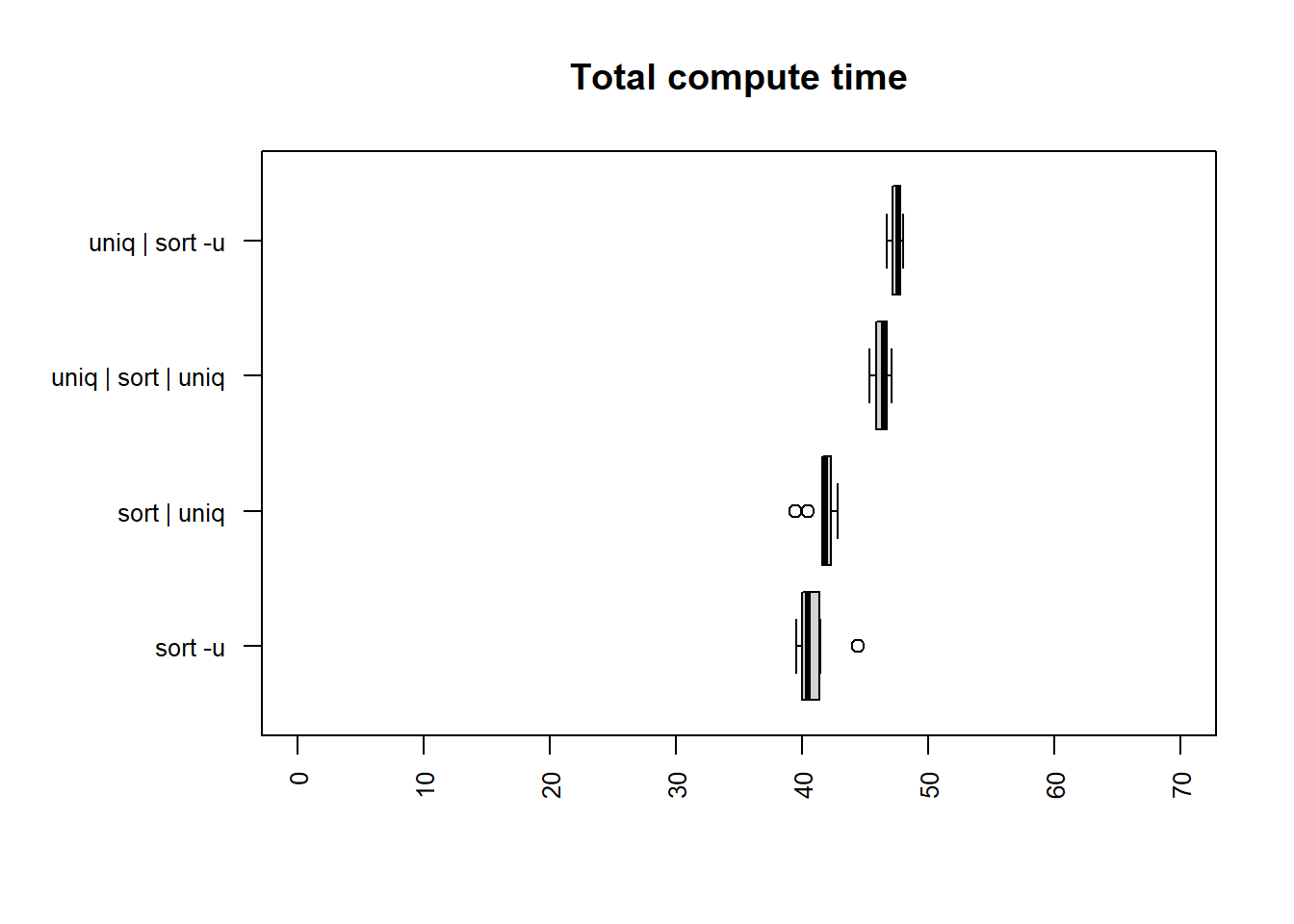
| method | mean |
|---|---|
| sort -u | 40.8204 |
| sort | uniq | 41.6528 |
| uniq | sort | uniq | 46.3127 |
| uniq | sort -u | 47.5305 |
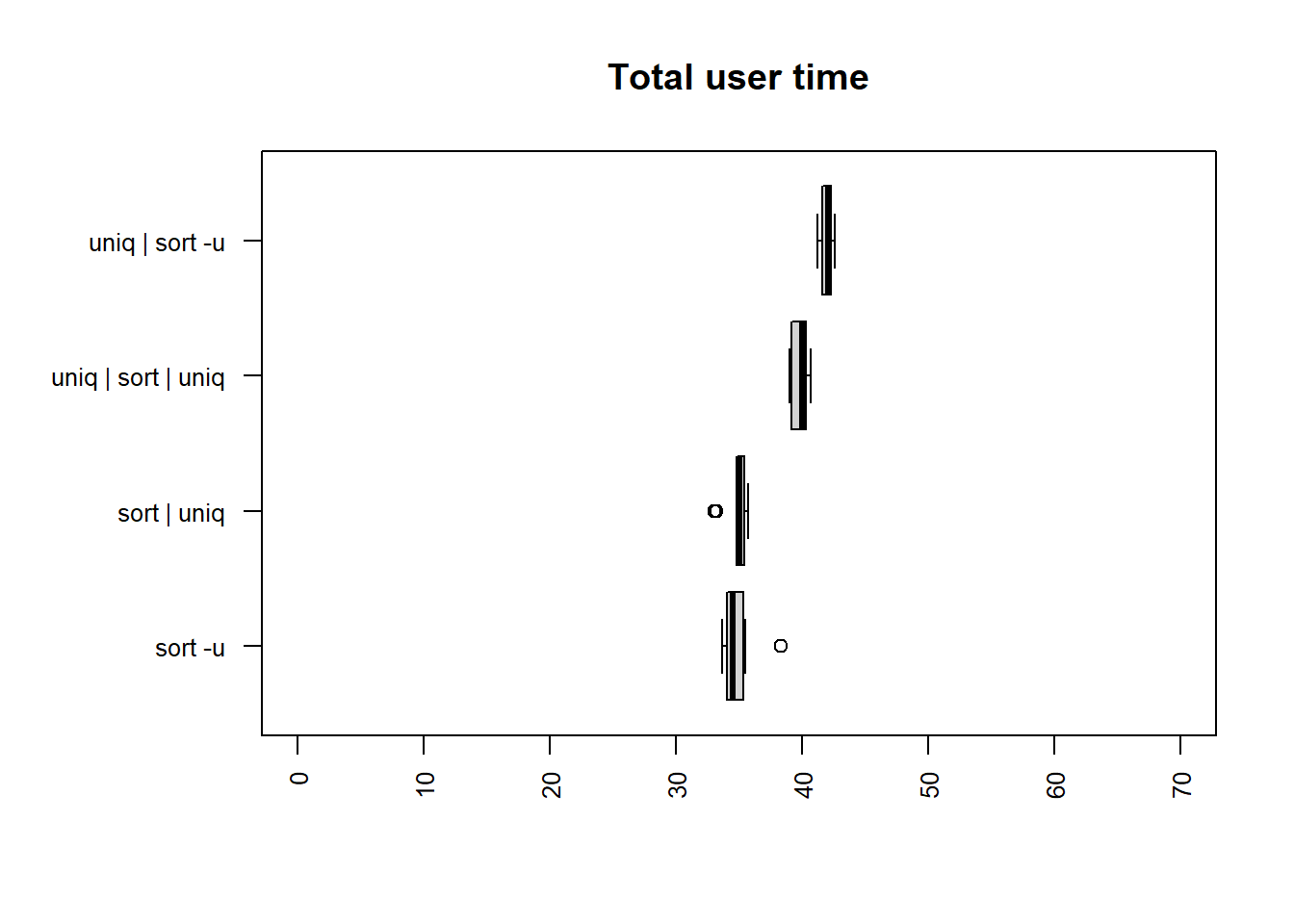
| method | mean |
|---|---|
| sort -u | 40.8204 |
| sort | uniq | 41.6528 |
| uniq | sort | uniq | 46.3127 |
| uniq | sort -u | 47.5305 |
The fastest times for both user and total time are
sort -u and sort | uniq. The calculation is
slower than for the license file in all cases apart from
the compute time for sort | uniq.
unique column
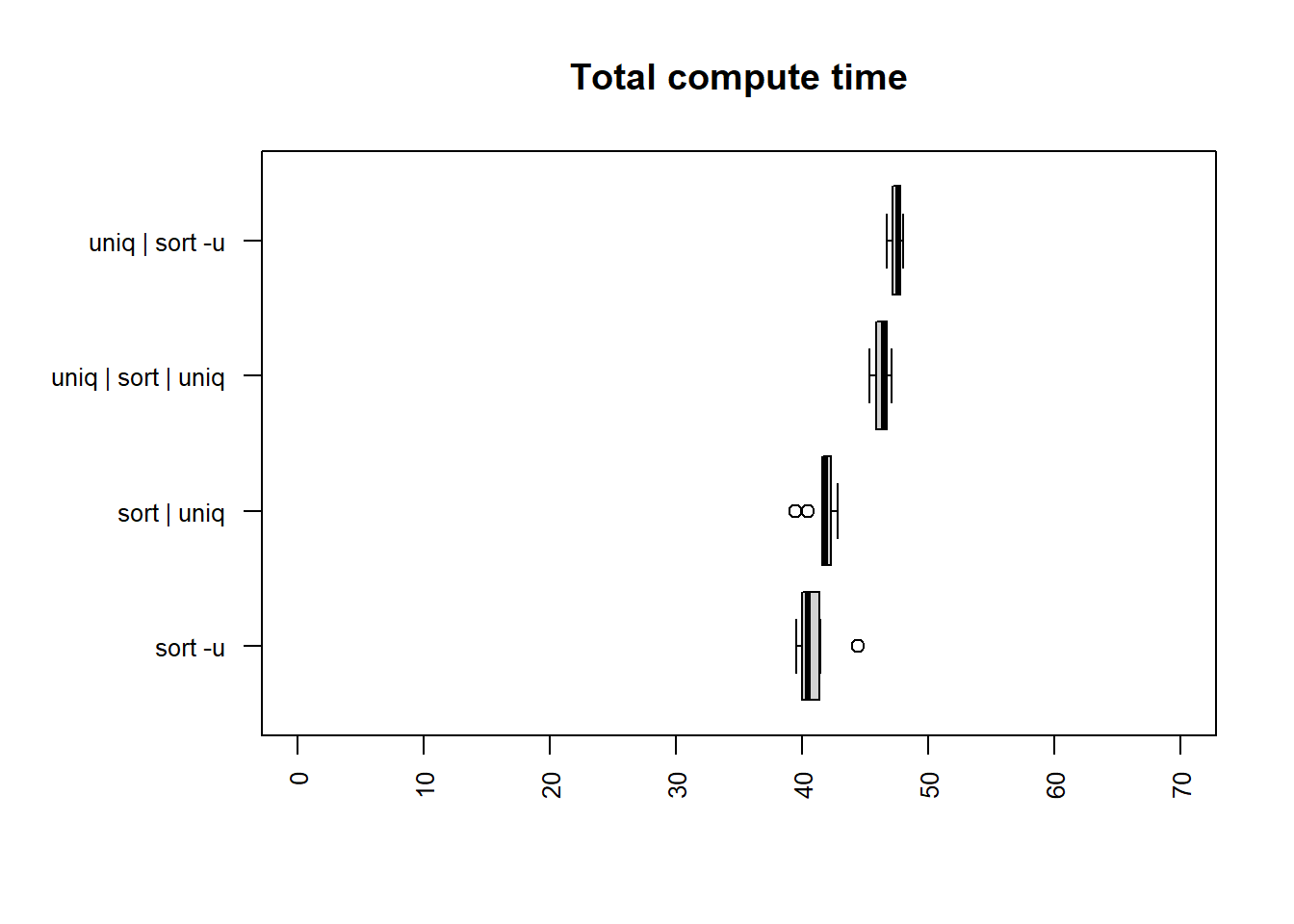
| method | mean |
|---|---|
| sort -u | 40.8204 |
| sort | uniq | 41.6528 |
| uniq | sort | uniq | 46.3127 |
| uniq | sort -u | 47.5305 |
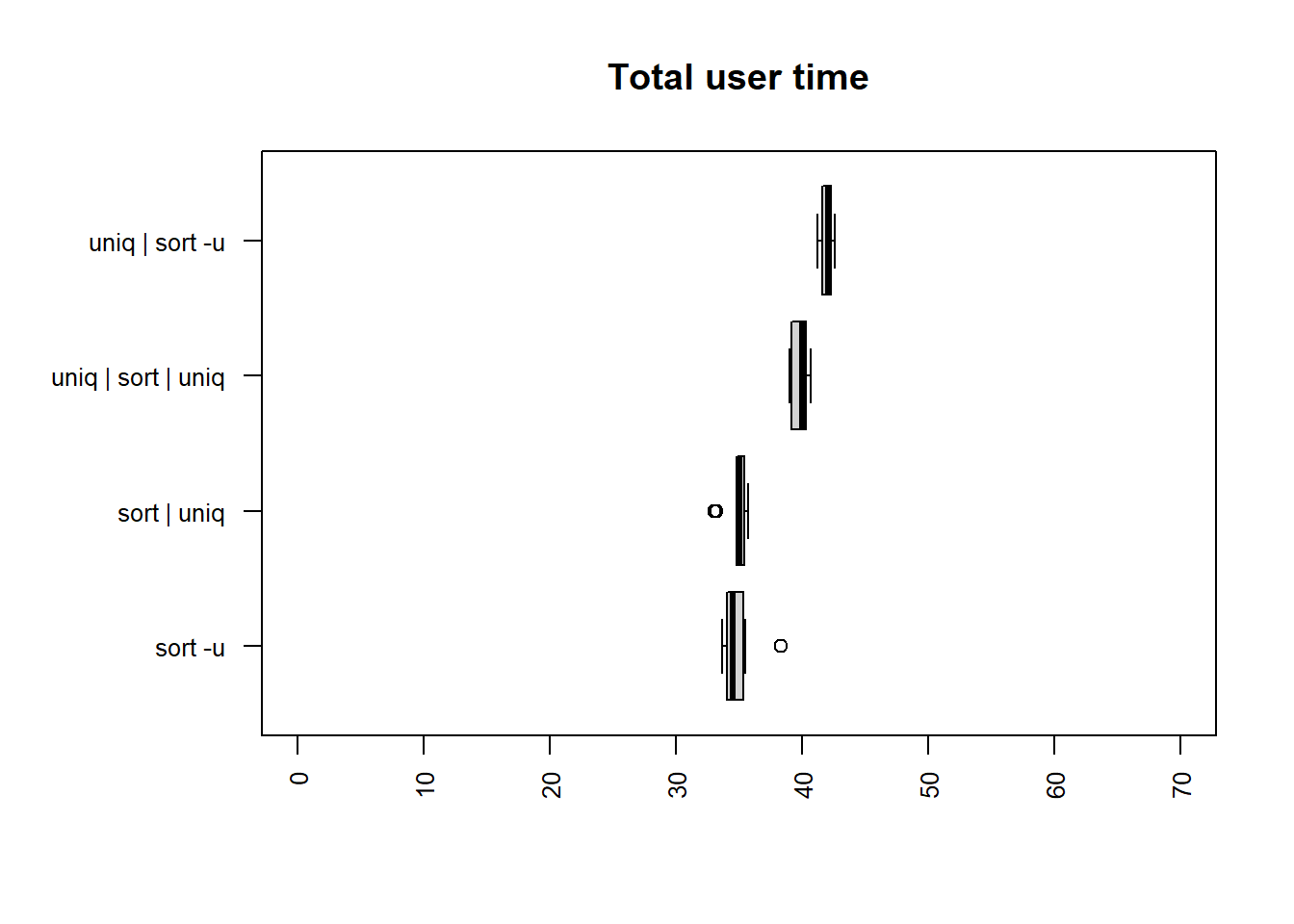
| method | mean |
|---|---|
| sort -u | 40.8204 |
| sort | uniq | 41.6528 |
| uniq | sort | uniq | 46.3127 |
| uniq | sort -u | 47.5305 |
The fastest times for both user and total time are
sort -u and sort | uniq, although all methods
are relatively closely matched.
alternating column
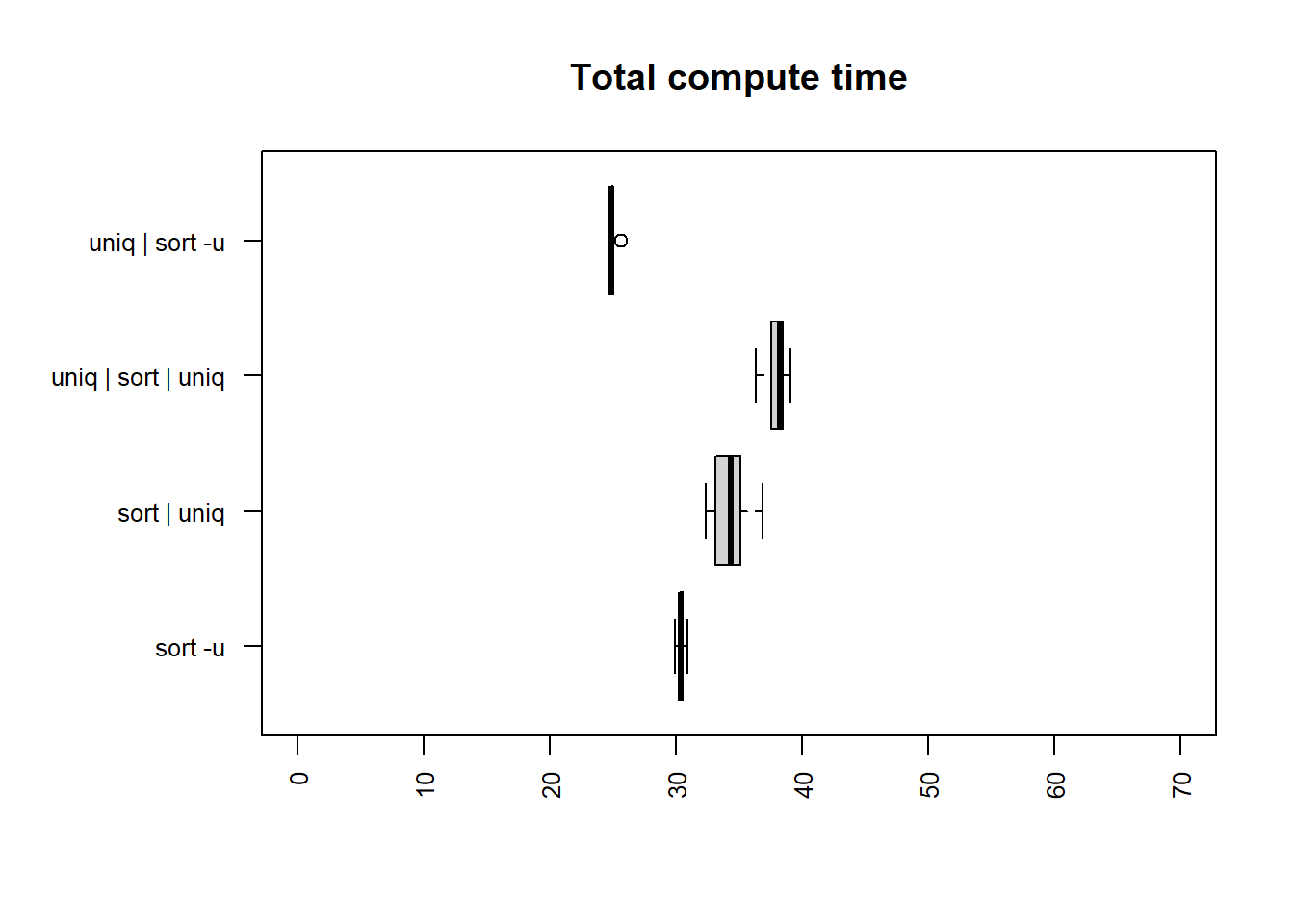
| method | mean |
|---|---|
| uniq | sort -u | 24.9655 |
| sort -u | 30.4055 |
| sort | uniq | 34.3696 |
| uniq | sort | uniq | 37.9967 |
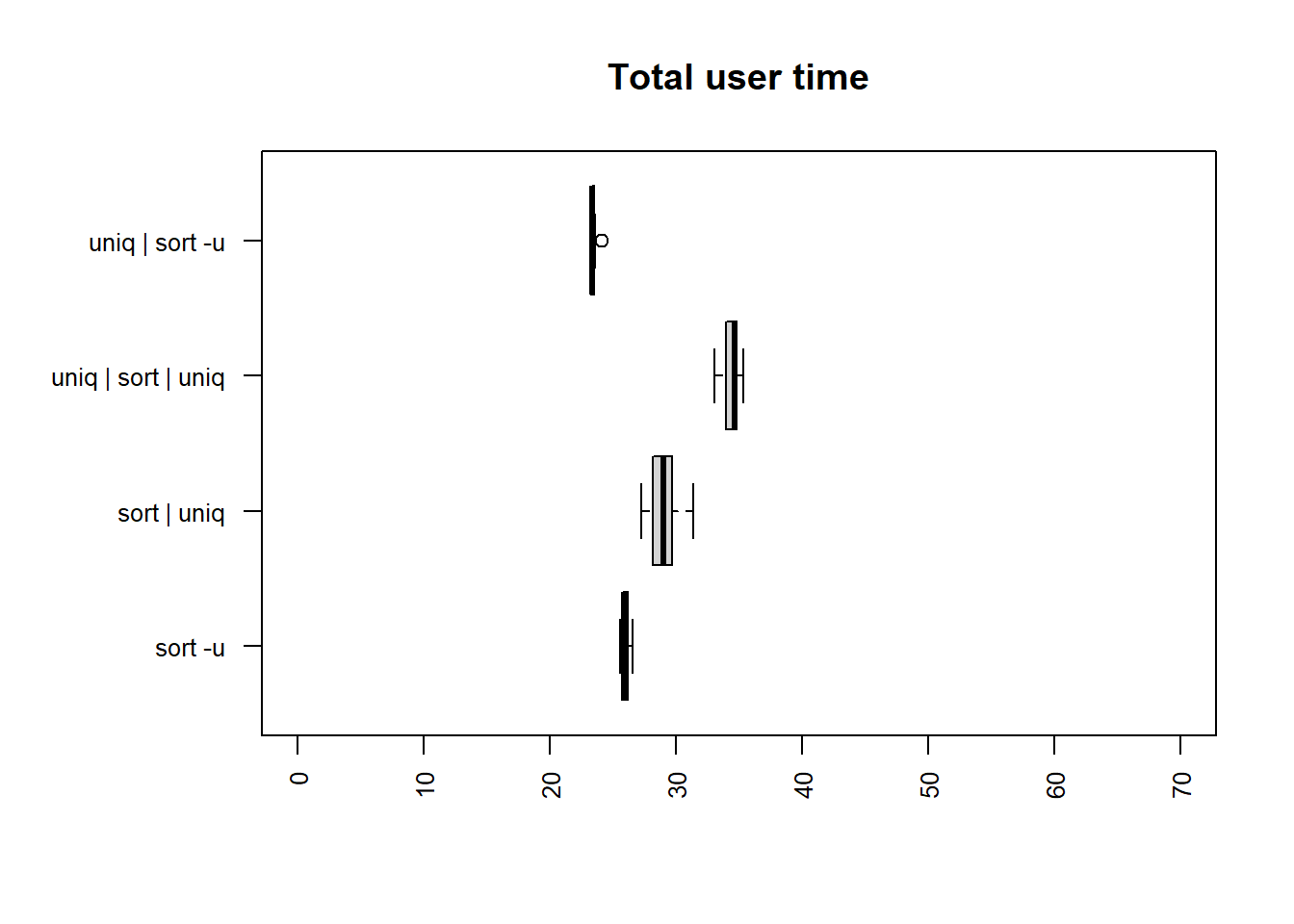
| method | mean |
|---|---|
| uniq | sort -u | 24.9655 |
| sort -u | 30.4055 |
| sort | uniq | 34.3696 |
| uniq | sort | uniq | 37.9967 |
The fastest times for both user and total time are
uniq | sort -u and sort -u.
same column
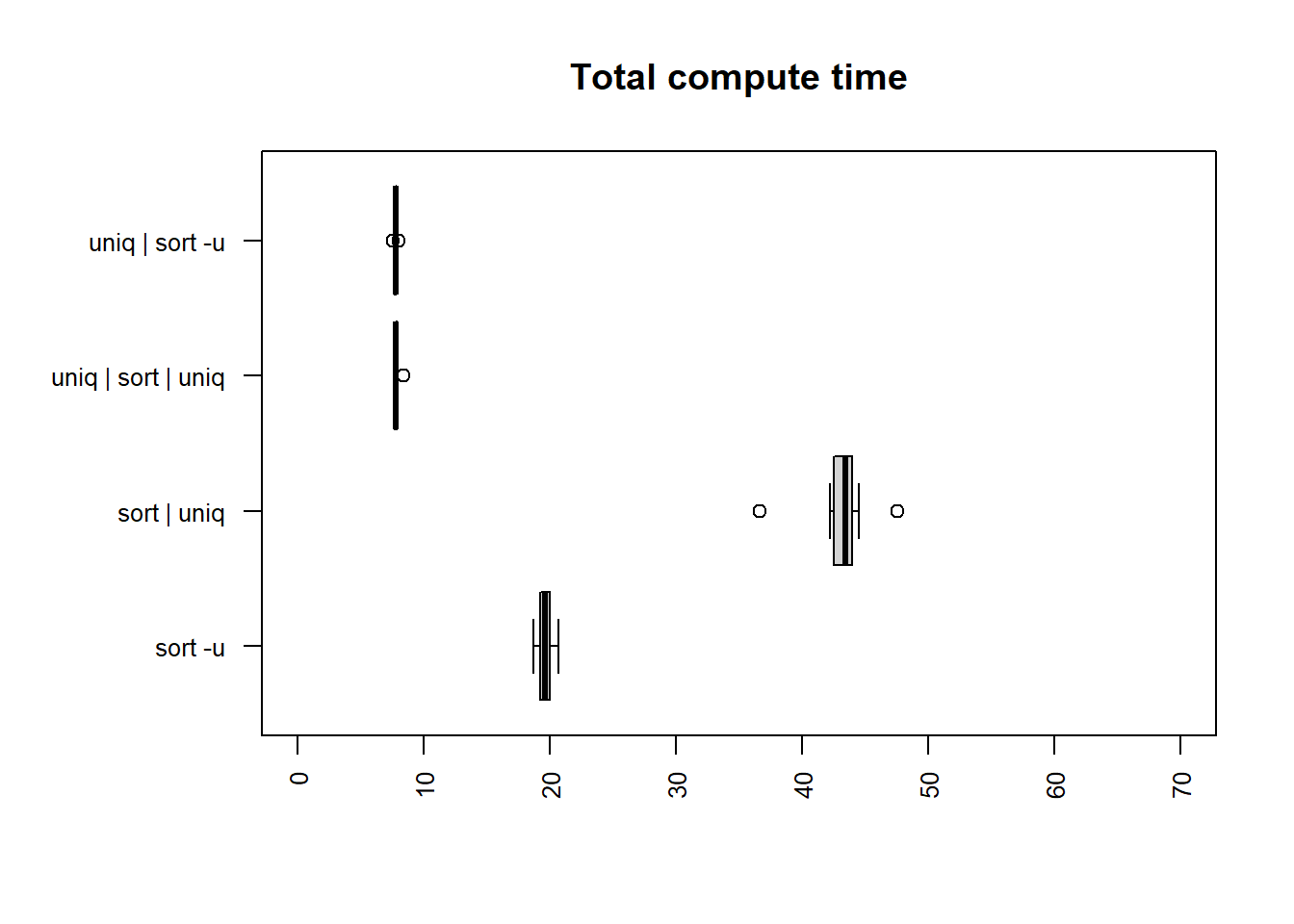
| method | mean |
|---|---|
| uniq | sort -u | 7.7747 |
| uniq | sort | uniq | 7.8341 |
| sort -u | 19.6351 |
| sort | uniq | 43.1565 |
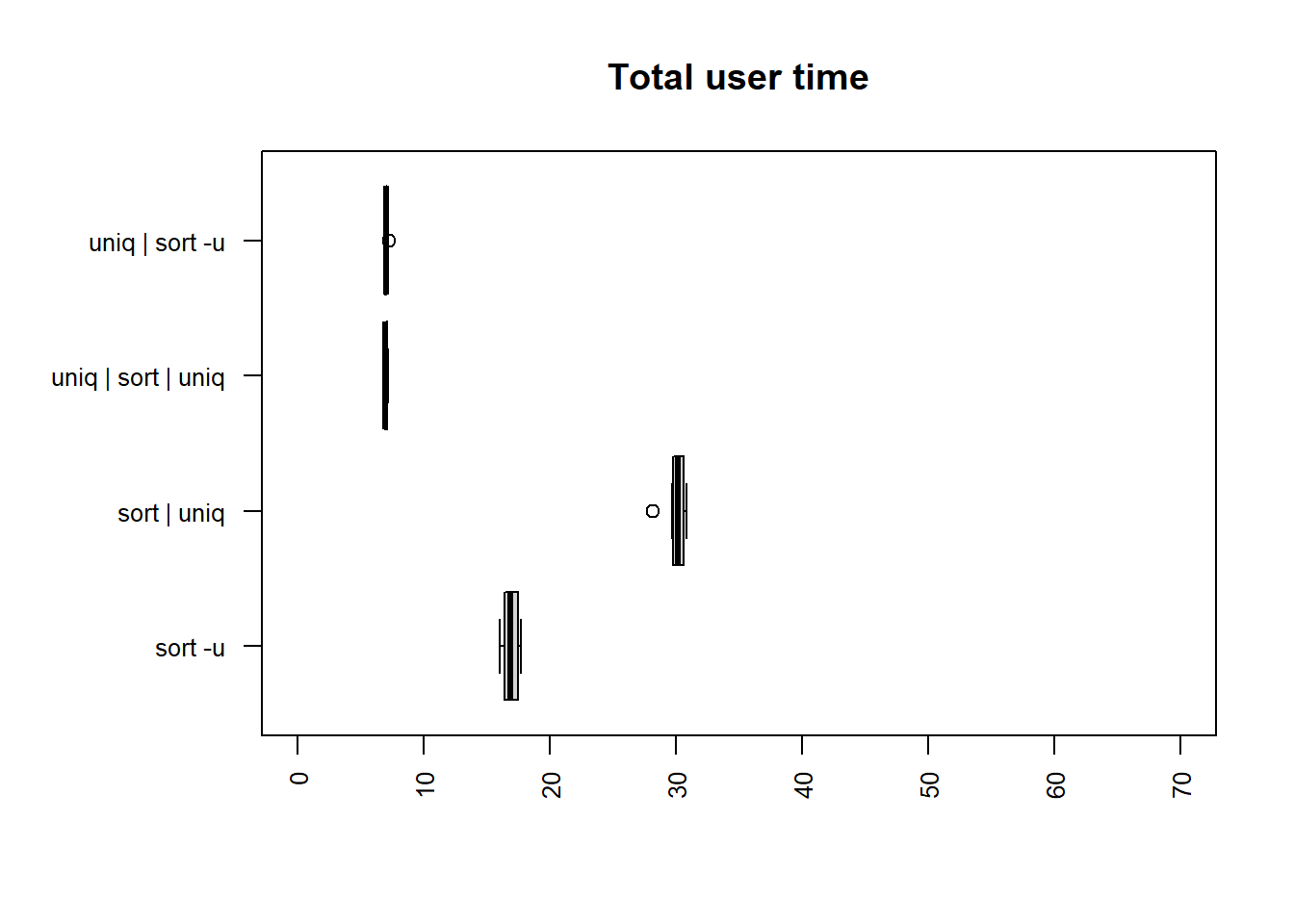
| method | mean |
|---|---|
| uniq | sort -u | 7.7747 |
| uniq | sort | uniq | 7.8341 |
| sort -u | 19.6351 |
| sort | uniq | 43.1565 |
The fastest times for both user and total time are
uniq | sort -u and uniq | sort | uniq, which
are similar in time taken and noticeably faster than the others.
Results by method
uniq | sort -u
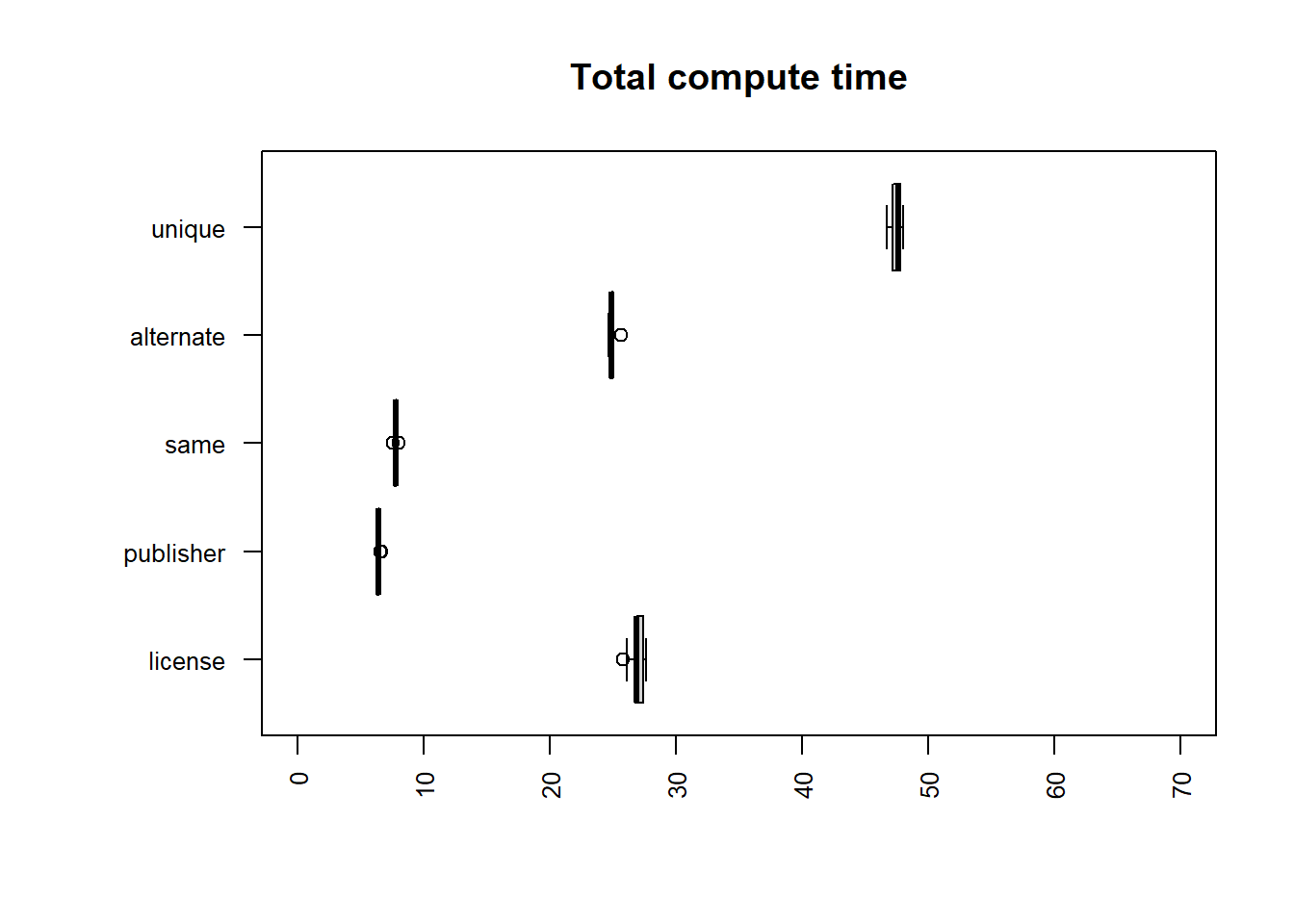
| file | mean |
|---|---|
| publisher | 6.4099 |
| same | 7.7747 |
| alternate | 24.9655 |
| license | 26.9240 |
| unique | 47.5305 |
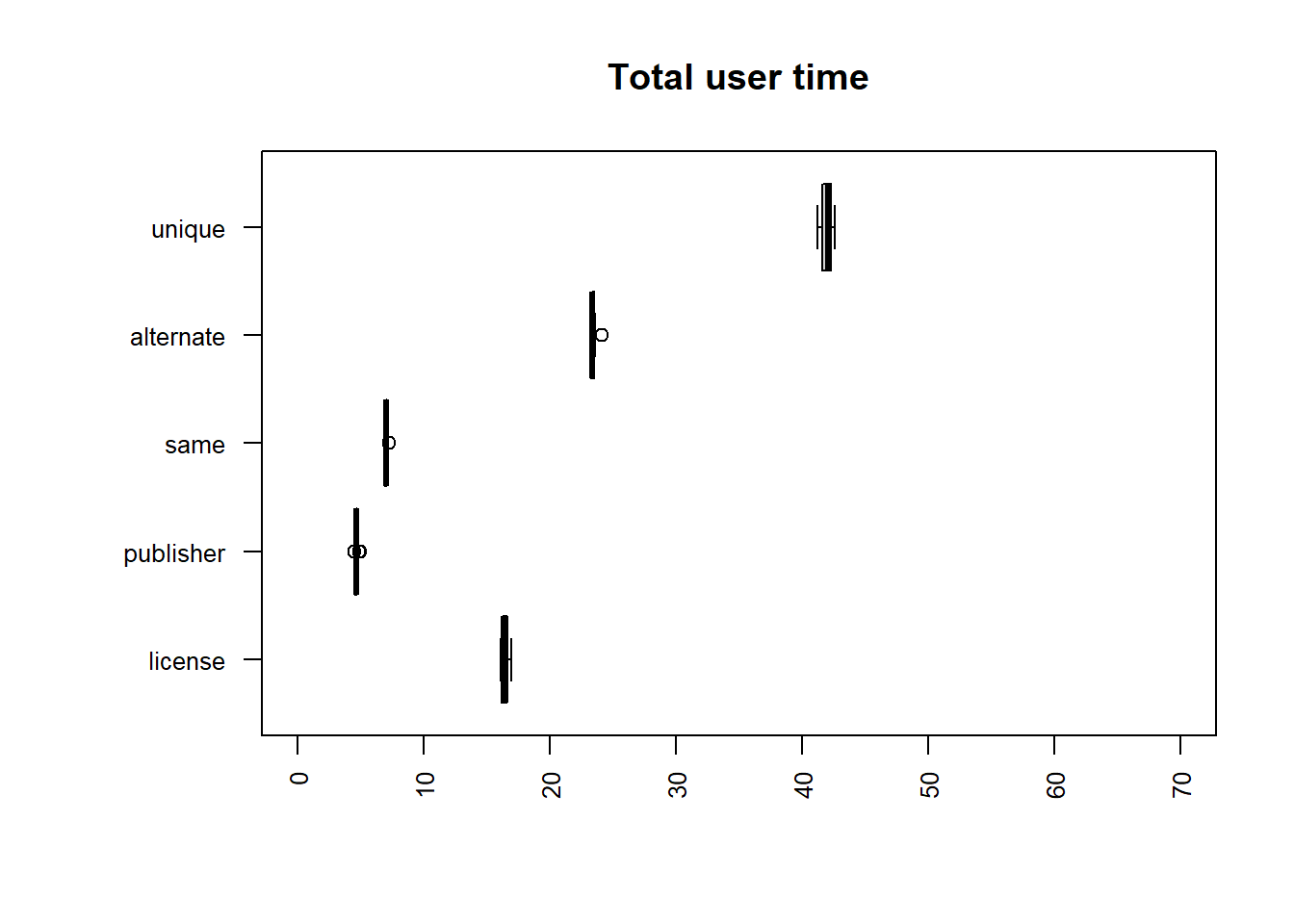
| file | mean |
|---|---|
| publisher | 6.4099 |
| same | 7.7747 |
| alternate | 24.9655 |
| license | 26.9240 |
| unique | 47.5305 |
sort | uniq
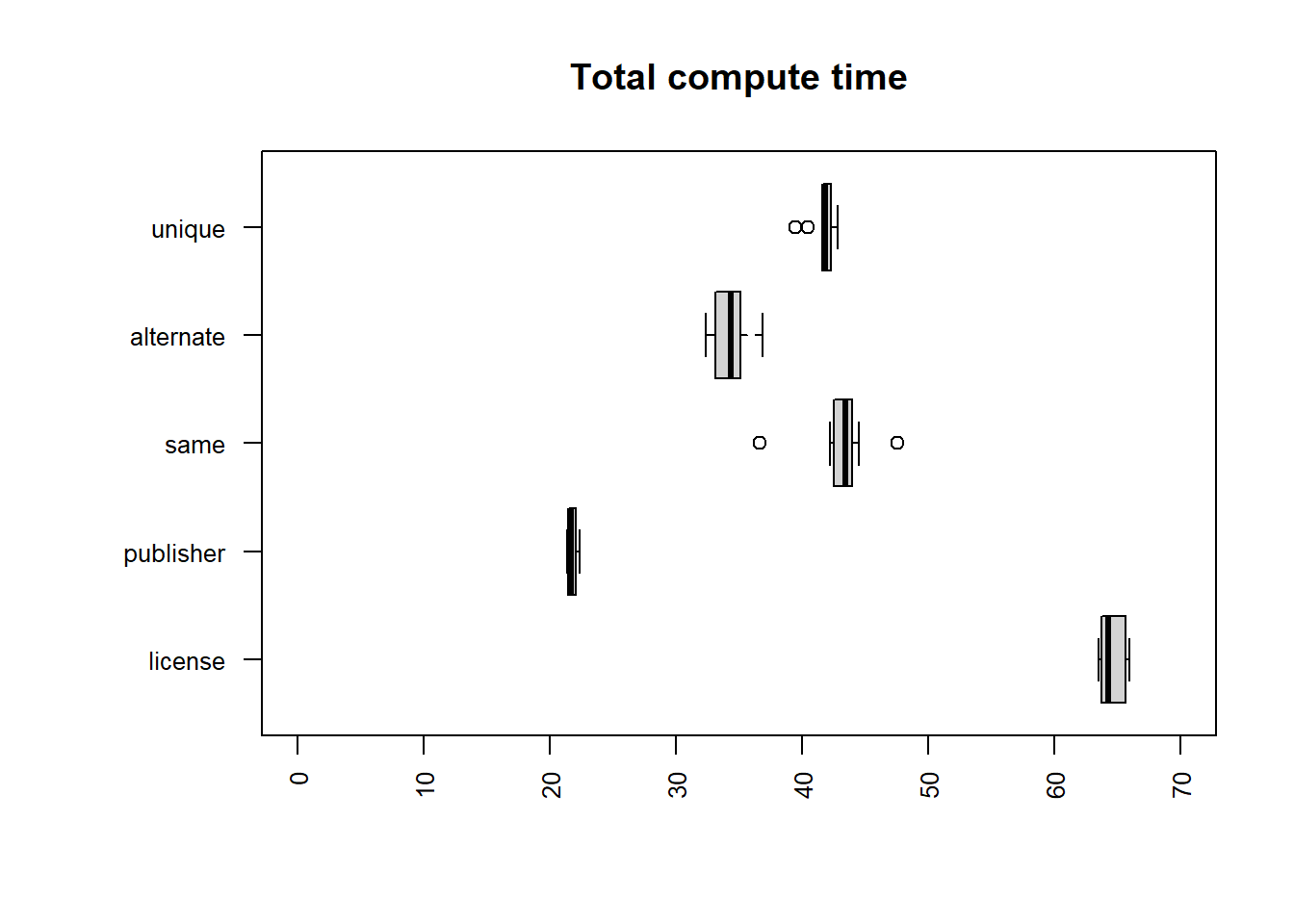
| file | mean |
|---|---|
| publisher | 21.7443 |
| alternate | 34.3696 |
| unique | 41.6528 |
| same | 43.1565 |
| license | 64.5948 |
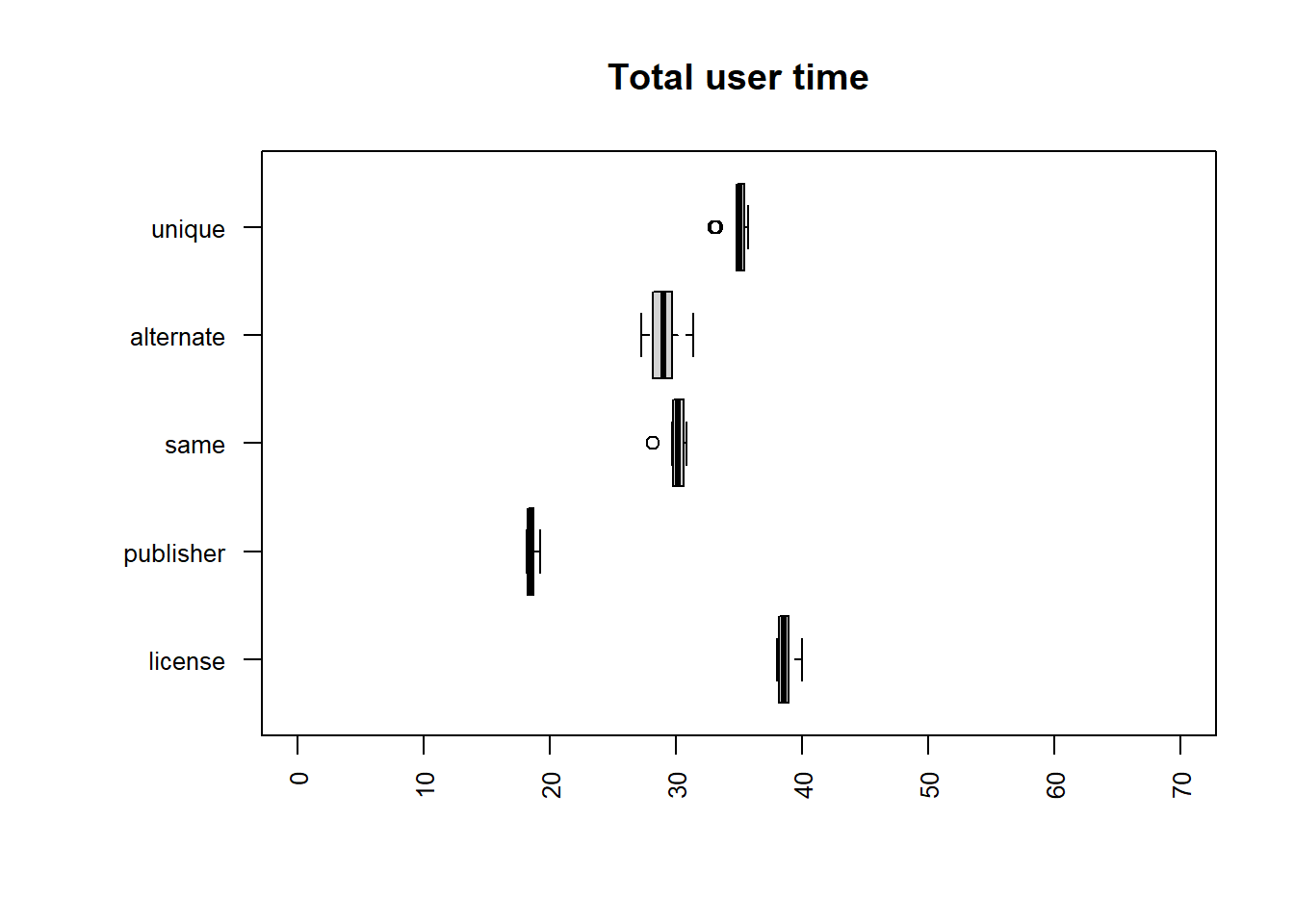
| file | mean |
|---|---|
| publisher | 21.7443 |
| alternate | 34.3696 |
| unique | 41.6528 |
| same | 43.1565 |
| license | 64.5948 |
uniq | sort | uniq
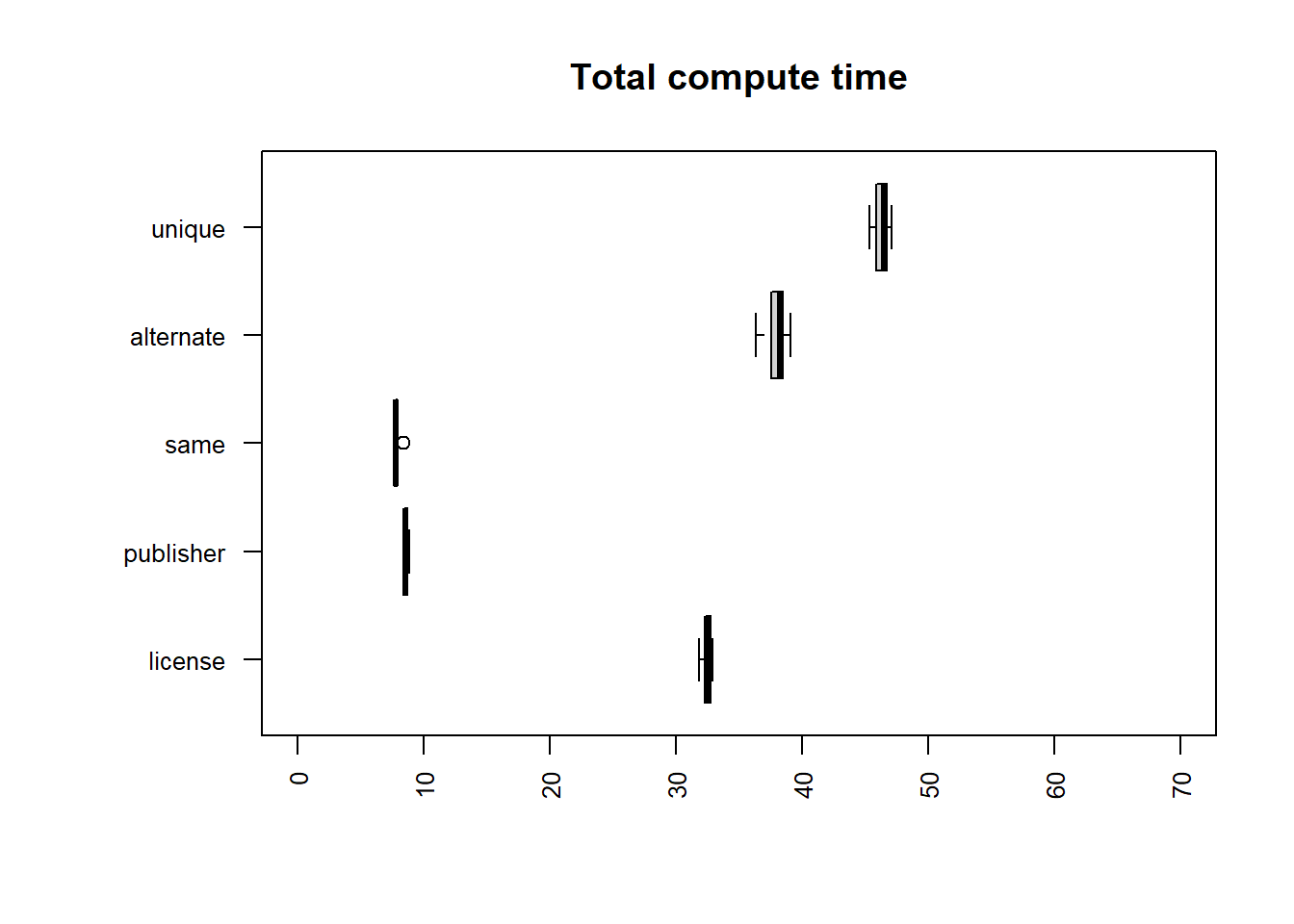
| file | mean |
|---|---|
| same | 7.8341 |
| publisher | 8.5796 |
| license | 32.5221 |
| alternate | 37.9967 |
| unique | 46.3127 |
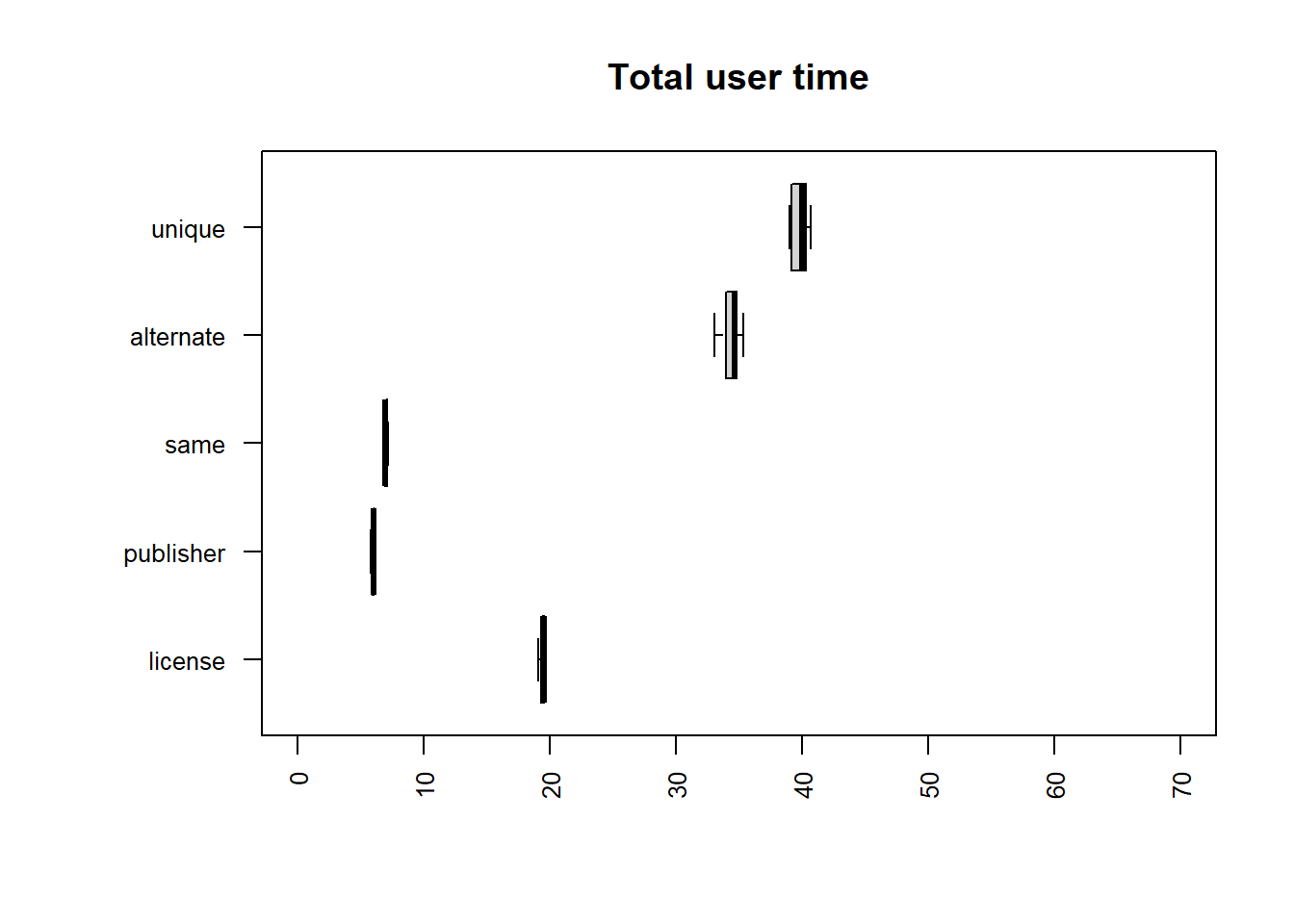
| file | mean |
|---|---|
| same | 7.8341 |
| publisher | 8.5796 |
| license | 32.5221 |
| alternate | 37.9967 |
| unique | 46.3127 |
uniq | sort -u
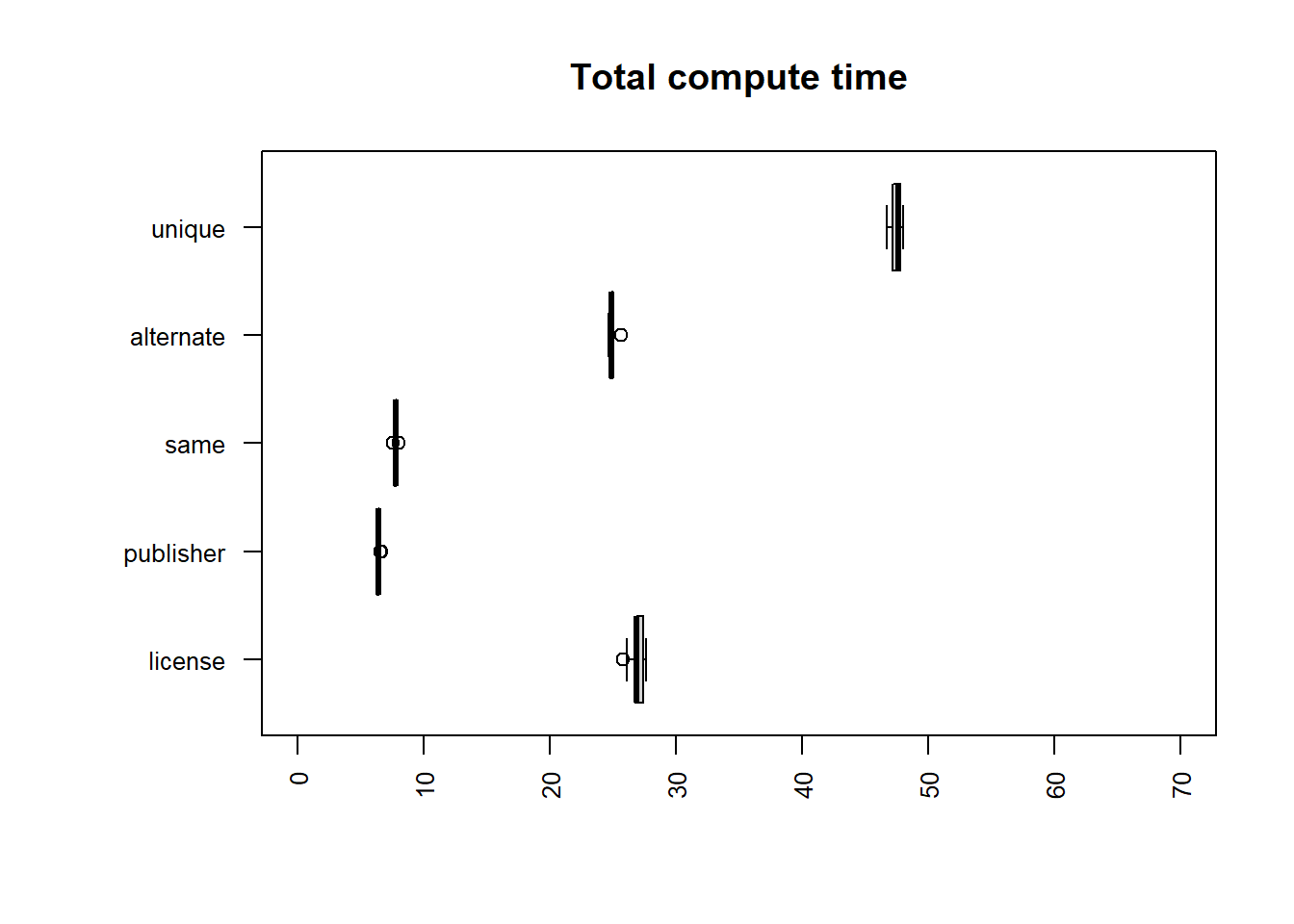
| file | mean |
|---|---|
| publisher | 6.4099 |
| same | 7.7747 |
| alternate | 24.9655 |
| license | 26.9240 |
| unique | 47.5305 |
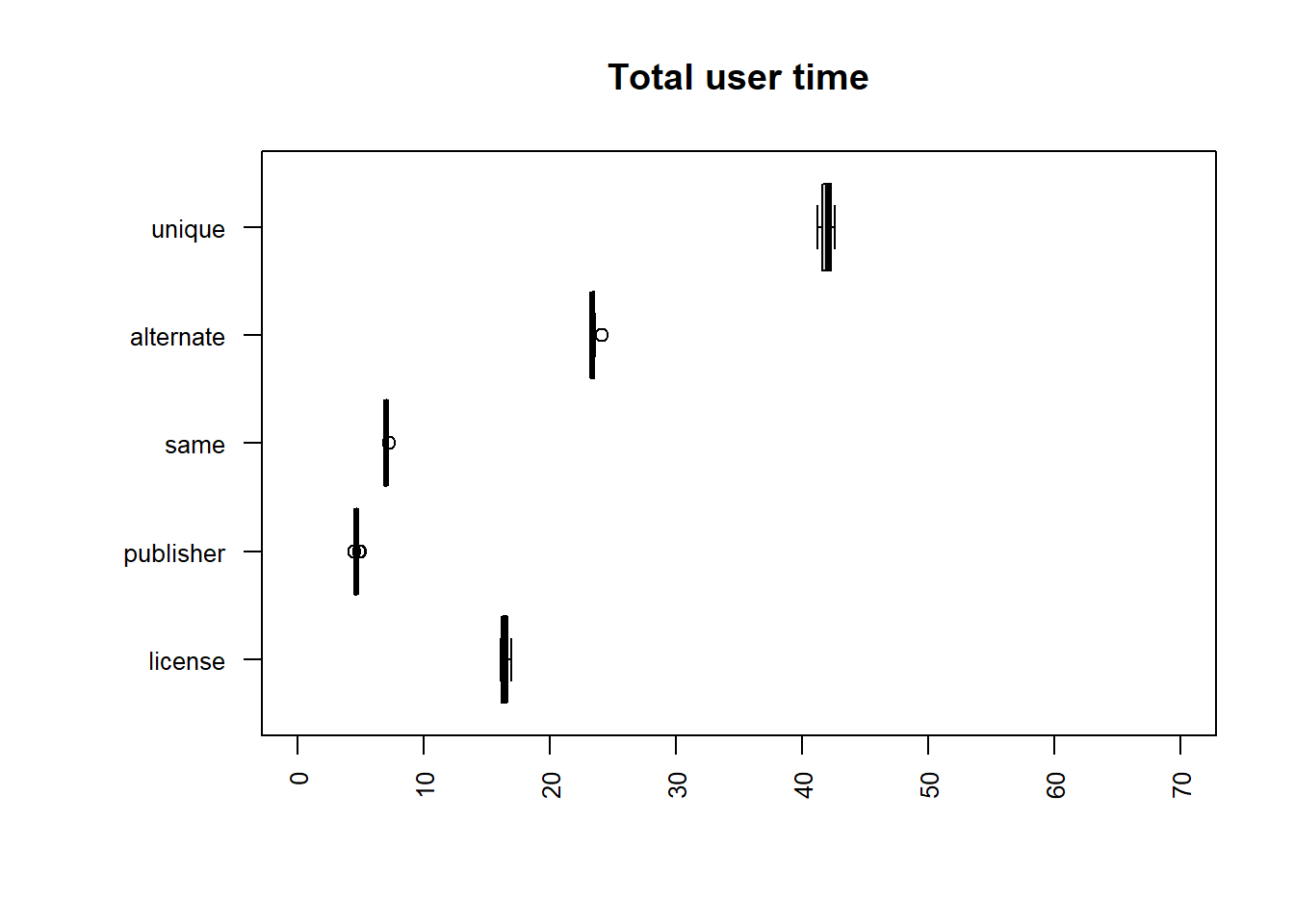
| file | mean |
|---|---|
| publisher | 6.4099 |
| same | 7.7747 |
| alternate | 24.9655 |
| license | 26.9240 |
| unique | 47.5305 |
Conclusions
For most real-world scenarios sort -u is probably a time
efficient choice. For very large datasets which are predominantly single
valued uniq | sort -u or uniq | sort | uniq
may be more efficient.
Acknowledgements
The virtual machine used for testing was provided by the Natural History Museum, London.